|
COVID-Align, one of the efforts to combat the COVID-19 pandemic, is an online alignment tool based on a profile HMM estimated using HMMER and about 2500 high-quality SARS-CoV-2 genomes. HMM-based sequence alignment methods are gaining popularity nowadays. The scoring parameters are implicitly trained, and thus, the alignment methods based on HMMs do not rely on explicit scoring systems, which makes them independent of the empirical scoring methods described in Section 2.3. The first two problems are about how likely a given HMM could generate a sequence and how the HMM could produce the corresponding alignment, and the third problem is about how to build the structure and estimate the parameters of the HMM based on given sequences, which could be either aligned or unaligned. According to the three problems that are interesting when using HMMs, the adoption of HMMs in sequence alignment has three corresponding issues: the scoring problem, the alignment problem, and the training problem. In terms of sequence alignment, an HMM is a statistical model that describes probability distribution over biological sequences. 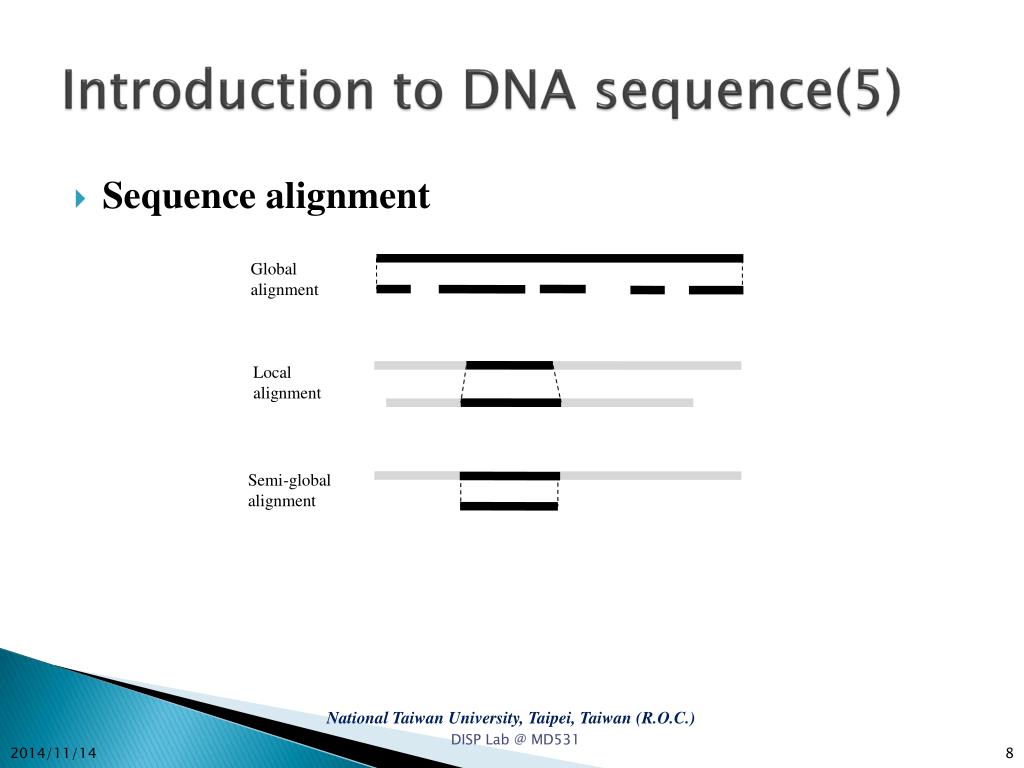
In addition, the hidden Markov model (HMM) is also widely utilized in sequence alignment tools, such as HHalign, which can perform high accurate profile HMM alignment. Although heuristic algorithms do not guarantee that there will be no poor results, ideal alignments can be achieved in many types of software because the sequences to be aligned are usually quite similar. For this, the most widely applied method is to limit the state transition and conduct the alignment in a smaller search space. Heuristic algorithms can be used to reduce the time and space cost incurred by dynamic programming. As a space-saving strategy of the dynamic programming algorithm, the Hirschberg algorithm is able to complete alignment by the space complexity of O( l) without any sacrifice of quality.Īn optimal solution for the pairwise sequence alignment of very long sequences is usually impossible to find in practice. Such overheads are acceptable for short sequences but not for sequences with more than several thousand sites. Time and space complexity of pairwise sequence alignment algorithms based on dynamic programming is O( l 1 l 2), where l 1 and l 2 are the lengths of the two sequences to be aligned. The algorithm usually consists of two steps: one is calculating the states of the dynamic programming matrix and the other is tracking back from the final state to the initial state of the dynamic programming matrix to obtain the solution of alignment. A commonly used global alignment algorithm is the Needleman–Wunsch algorithm, which has become the basic algorithm that is used in many types of multiple sequence alignment software. The former is to find and align the similar local region, and the latter is end-to-end alignment. Pairwise sequence alignment is the basis of multiple sequence alignment and mainly divided into local alignment and global alignment. In order to be distinct from the previous work, this review will try to present a general overview of the algorithms that prevail in this field and cover the work of the last several years. There have been several reviews for multiple sequence alignment, such as Refs. In this review, the pairwise sequence alignment algorithms and the corresponding scoring system, heuristic algorithms for multiple sequence alignment and their defects, and quality estimation methods used to test multiple sequence alignment software are reviewed. Finally, a character matrix with the same number of columns and rows that correspond to the number of the sequences is obtained. Sequence alignment software usually inserts gaps between the nucleotides or amino acid residues in the sequences, so that as many similar sites as possible can be aligned. Phylogenetics, comparative genomics, and protein structure and function prediction all depend on sequence alignment to look for conserved regions. 
Sequence alignment is one of the basic tasks in the processing of biological sequences, and the accuracy of alignment affects the subsequent analyses. The increasing number of sequences are challenging the automated sequence analysis procedures. The developments in sequencing technologies have enabled unprecedentedly fast sequencing speeds and large-scale sequencing capabilities.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
